library(tidyverse)
library(here)
library(vegan)
library(ggtext)
library(sysfonts)
subtidal <- read_csv(here("posts", "eds240-infographic", "data", "ucsb_fish_clean_obs_1999-2024.csv"))
taxon <- read_csv(here("posts", "eds240-infographic", "data", "PISCO_kelpforest_taxon_table.1.3.csv")) %>%
filter(campus == "UCSB")
fish_taxon <- left_join(subtidal, taxon, by = "classcode") %>%
select(year, site, side, zone, level, transect, classcode, fish_tl, depth, pctcnpy, count, Family, Genus, Species, common_name) %>%
janitor::clean_names()
fish_role <- read_csv(here("posts", "eds240-infographic", "data", "ucsb_fish_clean_tx_bm_1999-2024.csv")) %>%
filter(year > 2005) %>%
select(classcode, BroadTrophic, Targeted) %>%
distinct() %>%
group_by(classcode) %>%
summarise(BroadTrophic = paste(BroadTrophic, collapse = ", "),
Targeted = paste(Targeted, collapse = ", "),
.groups = 'drop') %>%
janitor::clean_names()
fish <- left_join(fish_taxon, fish_role, by = "classcode") %>%
janitor::clean_names()
fish_clean <- fish %>%
filter(!is.na(fish_tl)) %>%
group_by(site) %>%
mutate(total_years = length(unique(year))) %>% # Calculate number of years each site was surveyed
ungroup() %>%
mutate(island = case_when( # Assign sites to respective islands
str_detect(pattern = "SRI", string = site) ~ "Santa Rosa",
str_detect(pattern = "SCI", string = site) ~ "Santa Cruz",
str_detect(pattern = "SMI", string = site) ~ "San Miguel",
str_detect(pattern = "ANA", string = site) ~ "Anacapa",
TRUE ~ "remove"),
level = factor(level, levels = c("CAN", "MID", "BOT"))) %>% # Refactor survey levels
filter(island != "remove",
total_years >= 15,
year > 2005
) %>% # Select only sites with >=15 years of surveys
select(-total_years)
# Calculate canopy averages at transect level
kelp <- fish_clean %>%
group_by(site, side, transect, year) %>%
summarise(av_pctcnpy = round(mean(pctcnpy, na.rm = TRUE), digits = 0)) %>%
ungroup() %>%
mutate(cnpy_pct = case_when( # Bin percent canopy cover
av_pctcnpy <= 10 ~ "0-10",
av_pctcnpy <= 20 ~ "11-20",
av_pctcnpy <= 30 ~ "21-30",
av_pctcnpy <= 40 ~ "31-40",
av_pctcnpy <= 50 ~ "41-50",
av_pctcnpy <= 60 ~ "51-60",
av_pctcnpy <= 70 ~ "61-70",
av_pctcnpy <= 80 ~ "71-80",
av_pctcnpy <= 90 ~ "81-90",
av_pctcnpy > 90 ~ "91-100",
TRUE ~ NA,
)
) %>%
filter(!is.na(cnpy_pct))
fish_kelp <- left_join(fish_clean, kelp, by = join_by(site, side, transect, year)) # Attach bins to data
species_sums <- fish_kelp %>%
group_by(family) %>%
mutate(total_obs = n(),
total_counts = sum(count)) %>%
ungroup() %>%
mutate(fish_family = case_when(
total_counts >= 30000 ~ family,
TRUE ~ "Other"))
# Color palettes
plot_palette = c("#001806", "#7F6F2B", "#9A7408", "#D4C569", "#AA9D52", "#EBF4F2", "#B6E5F0", "#96B8A9", "#1E6848", "#004B5D", "#002B2B")
# fonts
font_add_google(name = "Spectral", family = "spectral")
font_add_google(name = "Inconsolata", family = "inconsolata")
species_sums$fish_family <- factor(species_sums$fish_family,
levels = c("Atherinopsidae", "Embiotocidae", "Labridae", "Pomacentridae", "Sebastidae", "Serranidae", "Other"),
labels = c("Baitfish", "Surfperch", "Wrasses", "Damselfish", "Rockfish", "Basses", "Other"))
level_labs <- c(CAN = "Canopy",
MID = "Midwater",
BOT = "Bottom")
unique_sp <- species_sums %>%
group_by(year, cnpy_pct, level, side, zone, site) %>%
summarize(transects = max(transect),
unique_species = length(unique(classcode))) %>%
ungroup() %>%
group_by(cnpy_pct, level) %>%
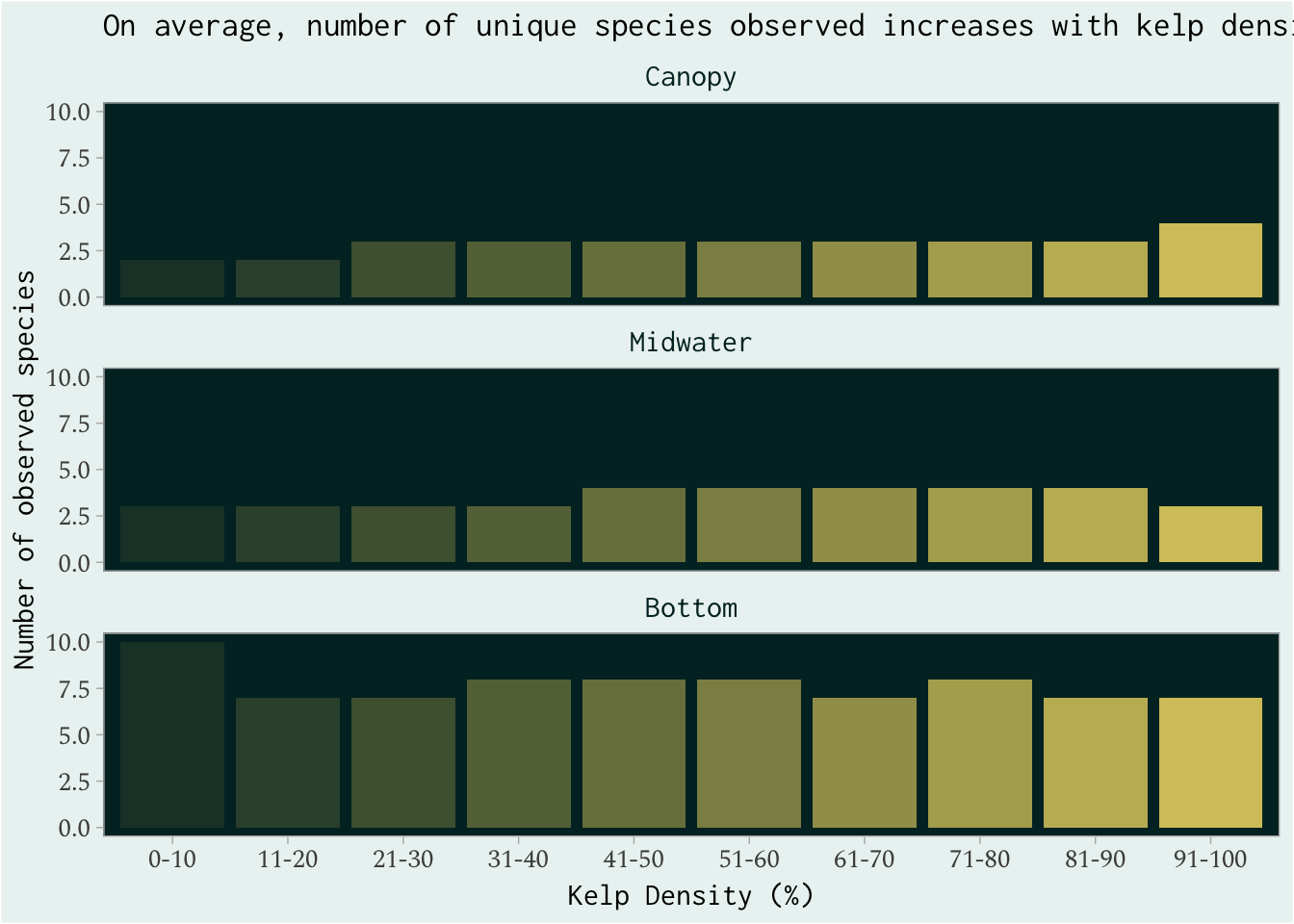
summarize(av_sp = round(mean(unique_species), 0))
ggplot(unique_sp) +
geom_col(aes(cnpy_pct, av_sp, alpha = cnpy_pct), fill = "#D4C569", color = NA, show.legend = FALSE) +
#geom_jitter(aes(cnpy_pct, unique_species, color = fish_family, size = total_fish), alpha = 0.9) +
#theme(legend.position = "none") +
scale_color_manual(values = fish) +
#scale_alpha_discrete(range = c(0.2,1)) +
facet_wrap(~level,
#scales = "free",
ncol = 1,
labeller = labeller(level = level_labs)) +
theme_light() +
theme(plot.title = element_markdown(family = "inconsolata"),
axis.title = element_markdown(family = "inconsolata", size = 12),
axis.text = element_markdown(family = "spectral", size = 10),
strip.text = element_markdown(family = "inconsolata", color = "#002B2B", size = 12),
panel.background = element_rect(fill = "#002B2B"),
panel.grid.major = element_line(linewidth = 0),
panel.grid.minor = element_line(linewidth = 0),
legend.background = element_rect(fill = NA),
legend.box.background = element_blank(),
plot.background = element_rect(fill = "#EBF4F2"),
strip.background = element_blank(),
strip.placement = "inside",
axis.title.x = element_text(margin = margin(t = 5, r = 0, b = 0, l = 0, unit = "pt")),) +
labs(x = "Kelp Density (%)",
y = "Number of observed species",
#color = "Family",
#size = "Total observed fish",
title = "On average, number of unique species observed increases \nwith kelp density and depth")